Working With Your Sequence
In this section, you will learn how to:
- start working with a sequence by adding it;
- edit sequence: copy and paste;
- edit sequence: add a feature (annotate);
- save and export your file.
Adding a Sequence
To add a sequence, you have two options: uploading the existing one or creating one from scratch.
A) Uploading the existing file. Mendelgen supports .gb, .gbk, .dna, .fasta, .txt sequence formats.
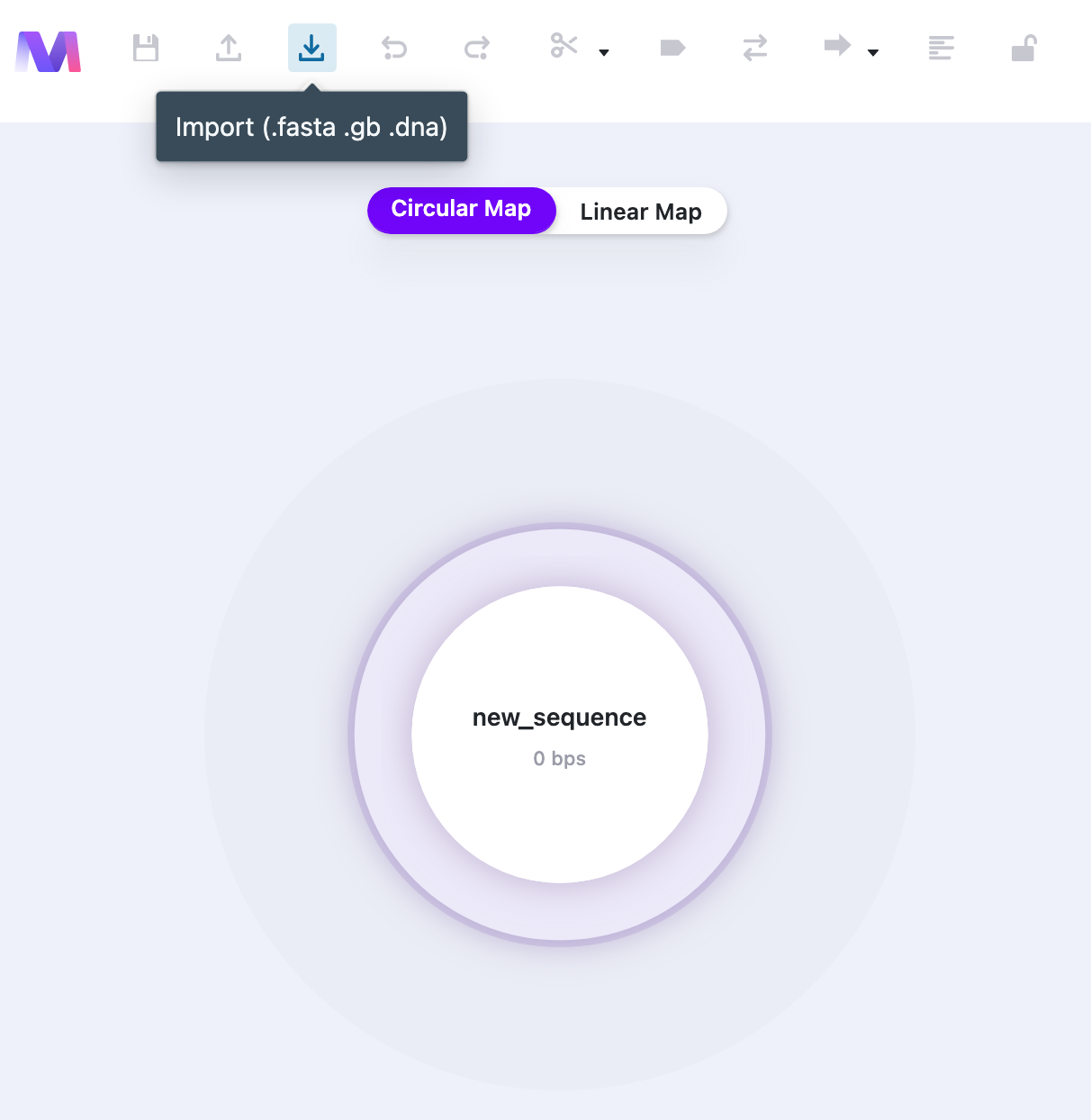
On the Dashboard, select “New Vector” and then click the “Import” button. The File Explorer pops up and you will be able to select a file from your computer. All the annotations (if available) are imported automatically.
B) Adding manually. To manually add the sequence, copy it from the source and paste it into the window (“Insert Sequence Here”).
- On the Dashboard, select “New Vector”:
- Paste the sequence into the field (“Insert Sequence Here”):
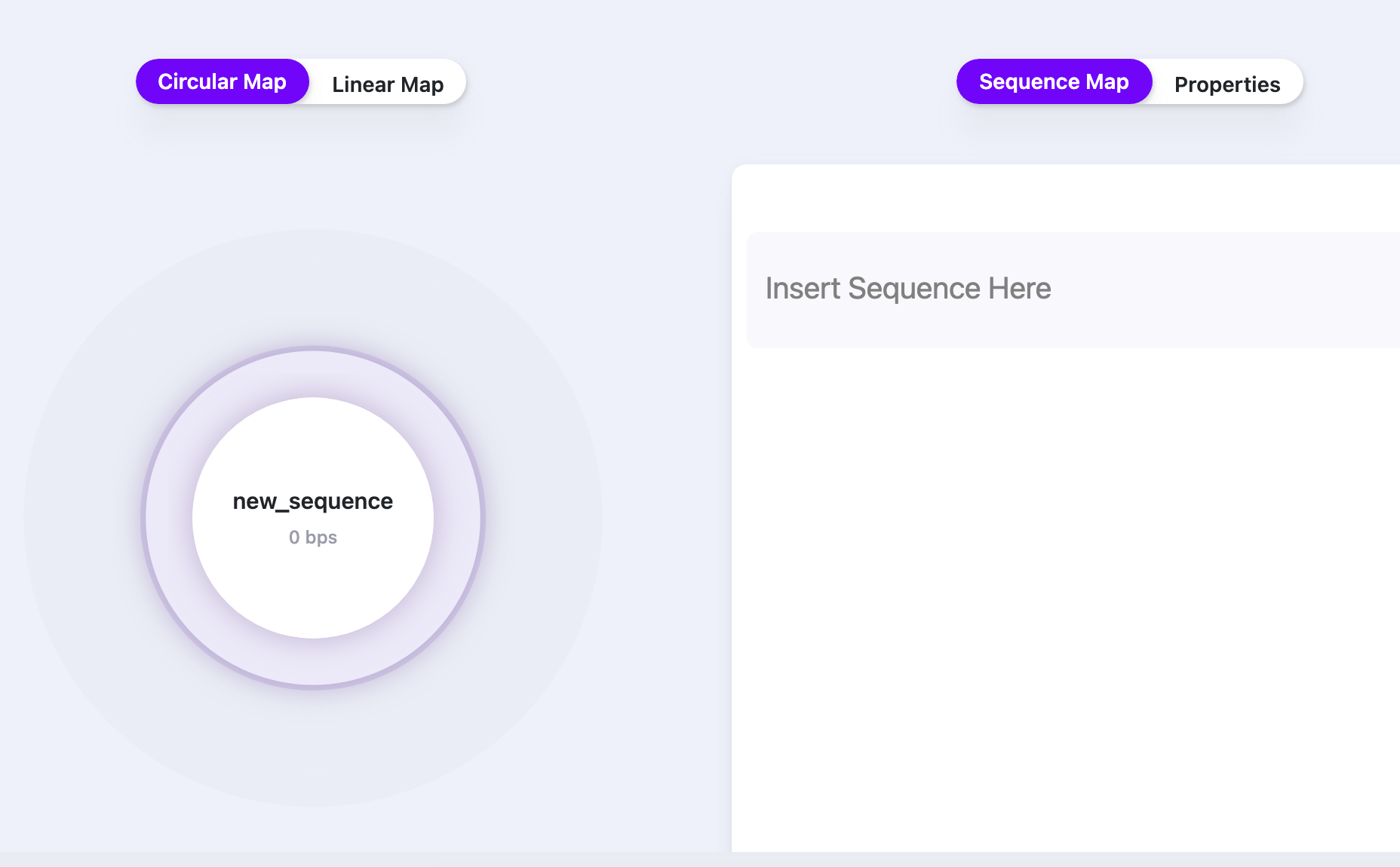
Editing a Sequence
Once you have added your sequence, there are several ways you can work with it. You can copy parts of the sequence and add them. You can annotate the sequence and group these features. You are able to print the sequence as well.
Copy/Paste a Section of DNA
To copy a section of the DNA sequence, simply place a caret at the beginning (before the first base of the desired sequence) and drag it to the end of the sequence. Press ⌘/Ctrl+C or use Right-click and select “Copy”.
You may use the “Copy” function in multiple ways. Copy Options allows you to copy the sequence with all metadata, to use with Mendelgen.
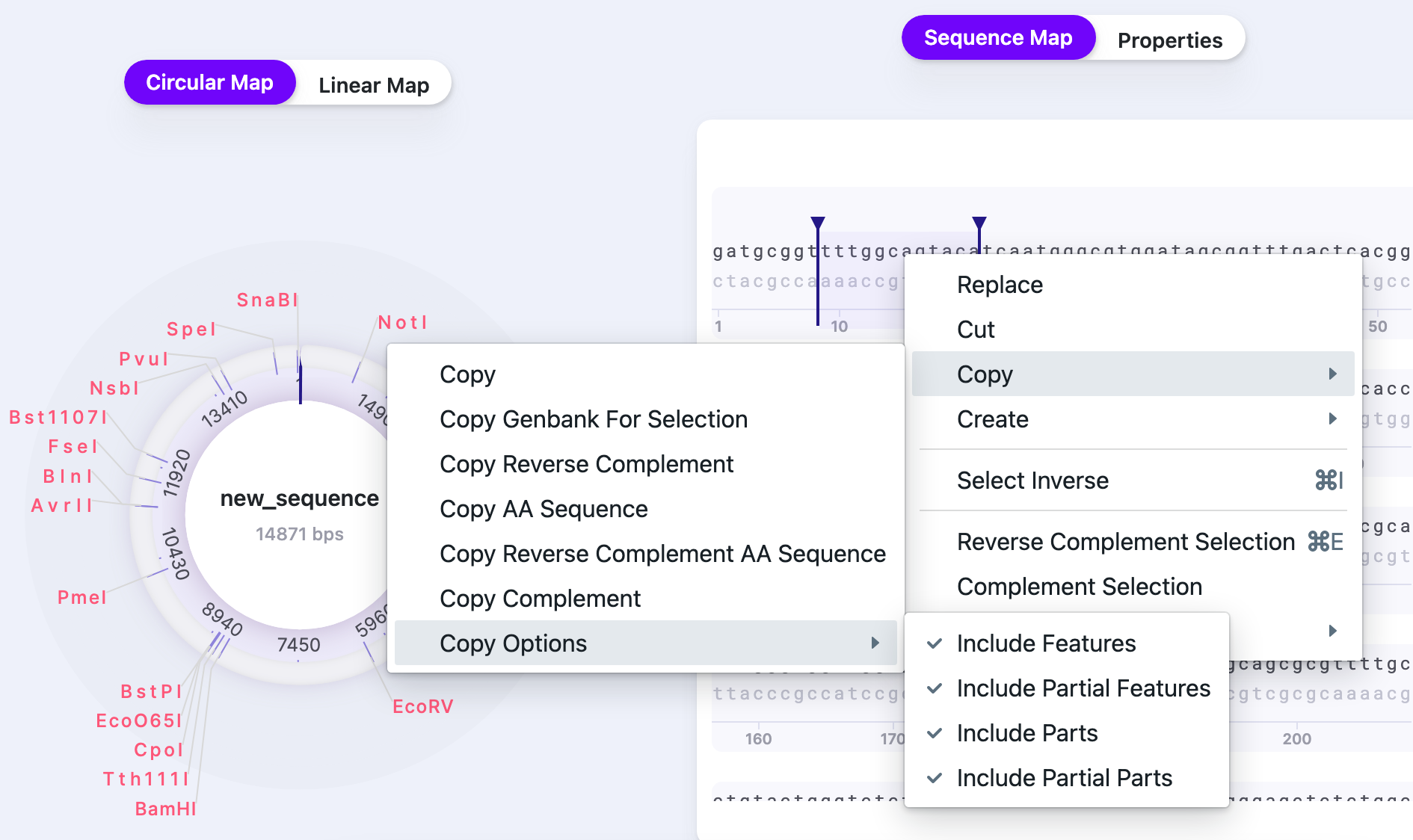
Example: with selecting the sequence: gatgcggttttggcagtacat, the copying options provide the following results:
| Copy | gatgcggttttggcagtacat |
| Copy GenBank for Selection | LOCUS new_sequence_partial 21 bp DNA linear DD-MM-YYYY COMMENT teselagen_unique_id: 5adf735aa1811801e17d8aac ORIGIN 1 gatgcggttt tggcagtaca t // |
| Copy Reverse Complement | atgtactgccaaaaccgcatc |
| Copy AA Sequence | DAVLAVH |
| Copy Reverse Complement AA Sequence | MYCQNRI |
| Copy Complement | ctacgccaaaaccgtcatgta |
To paste a section of the DNA sequence, place a caret at the desired location. Press ⌘/Ctrl+V or use Right-click and select “Insert”. You can see the position of the caret on the Circular or Linear map on the left in real-time.
Adding a Feature or a Part
The Features are the annotations for the major parts of your sequence, such as Primers, Promoters, RBS, Antibiotic Resistance Gene.
The Parts are comments on the sequence such as “overhang” or “linker”.
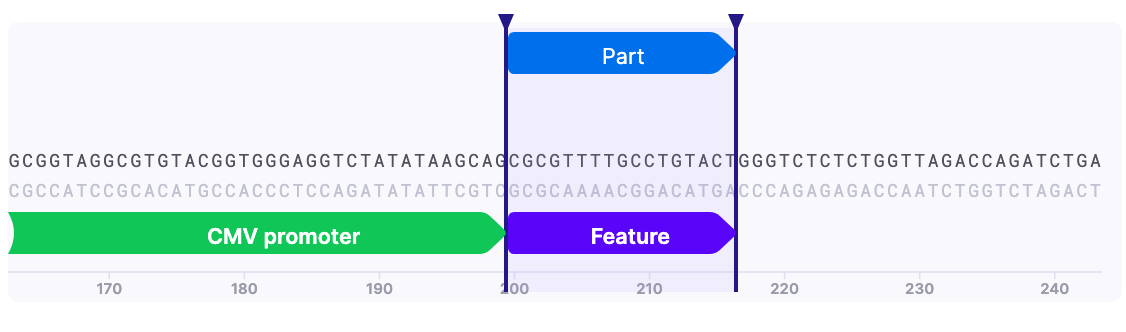
To add a feature:
1) Select the sequence, which you want to annotate
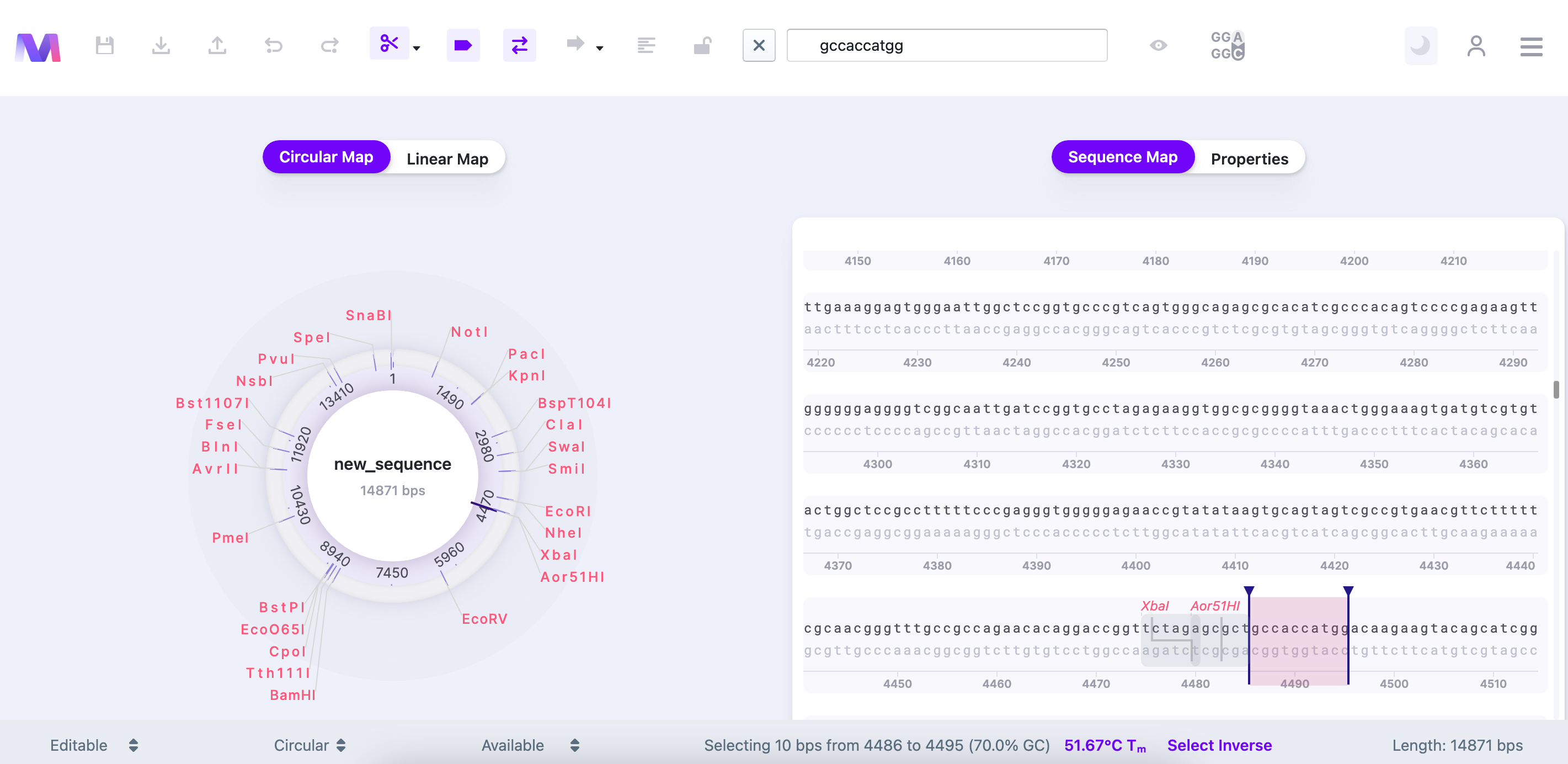
2) Right-click on the highlighted sequence OR ⌘/Ctrl+K
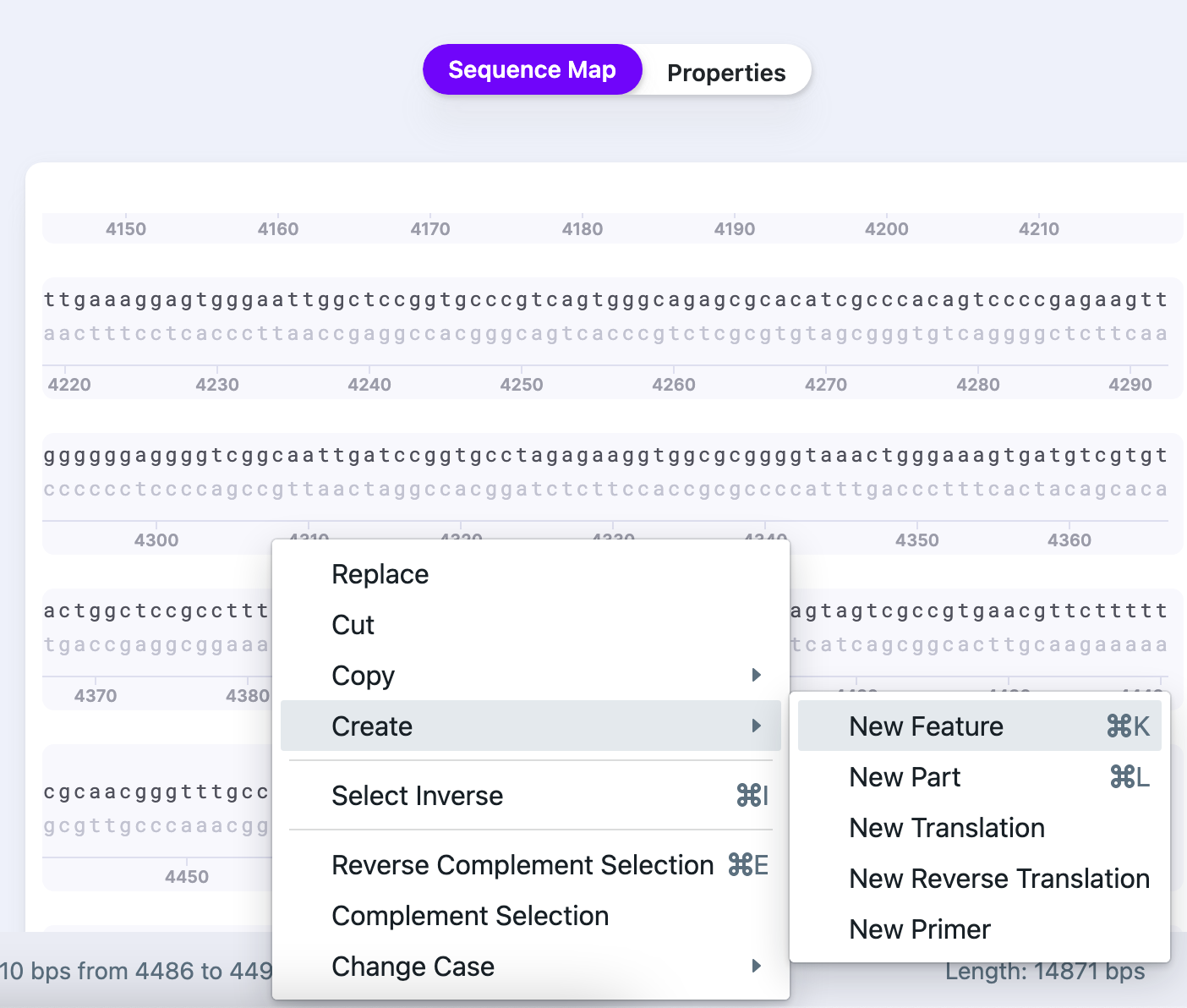
3) Select “New Feature”.
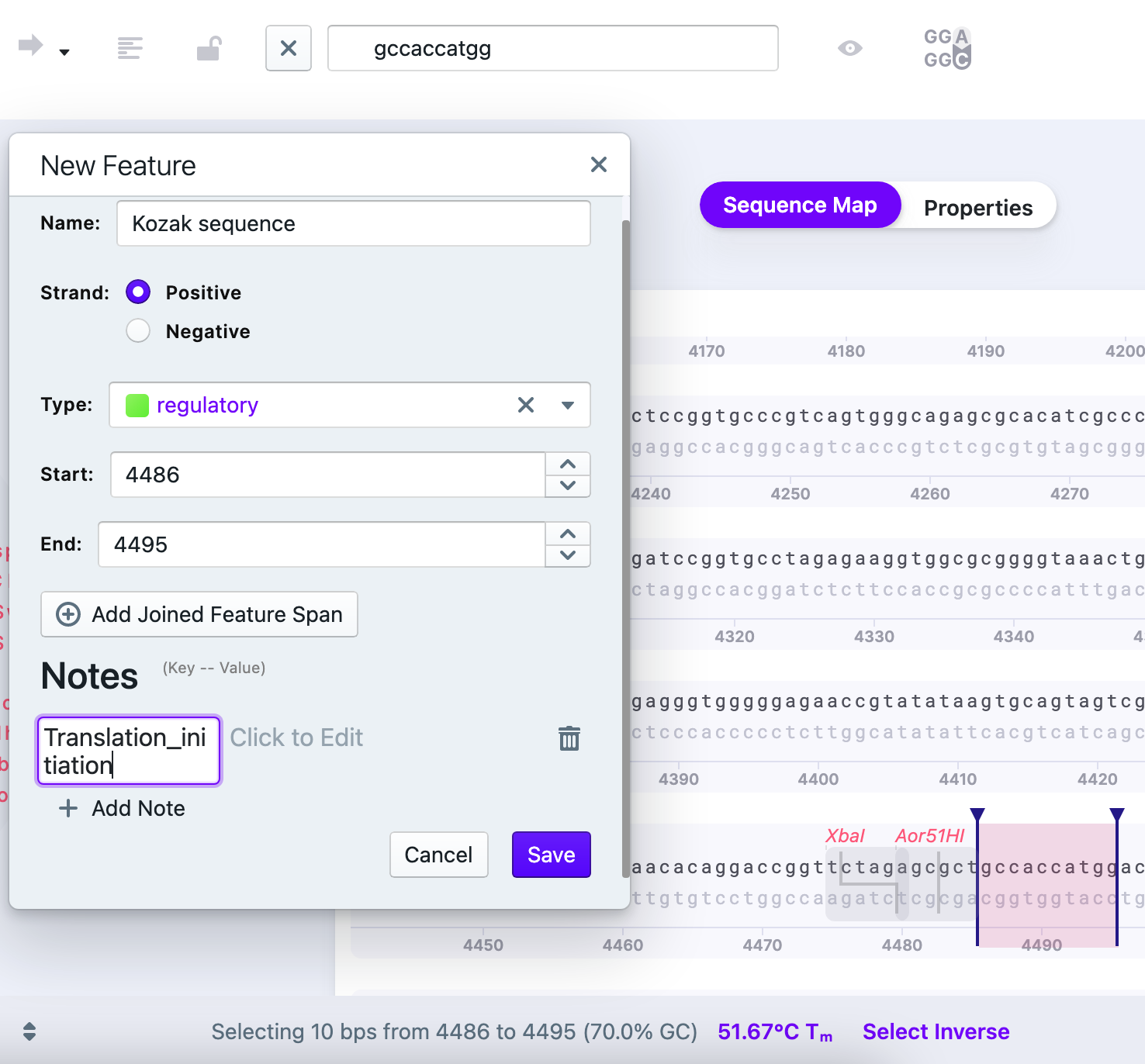
4) Write the name and confirm the strand. You can pick the type of feature as well. You may add several Notes about the selected part of the sequence. You also have an option of joined feature span. Press “Save”. The saved feature will be now displayed.
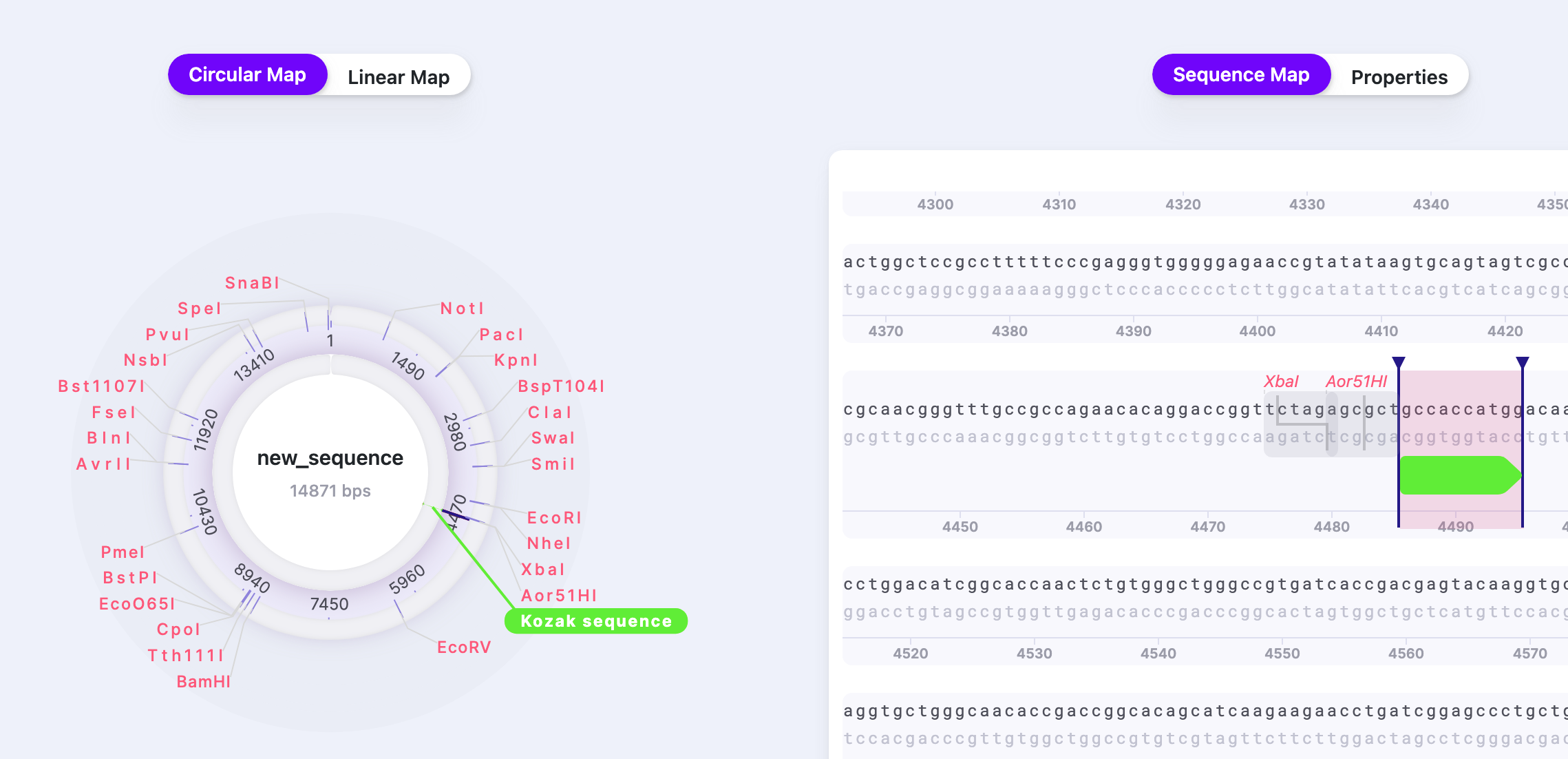
To add a part:
1) Select the sequence and press ⌘/Ctrl + L or select Right-click –> Create –> New Part.
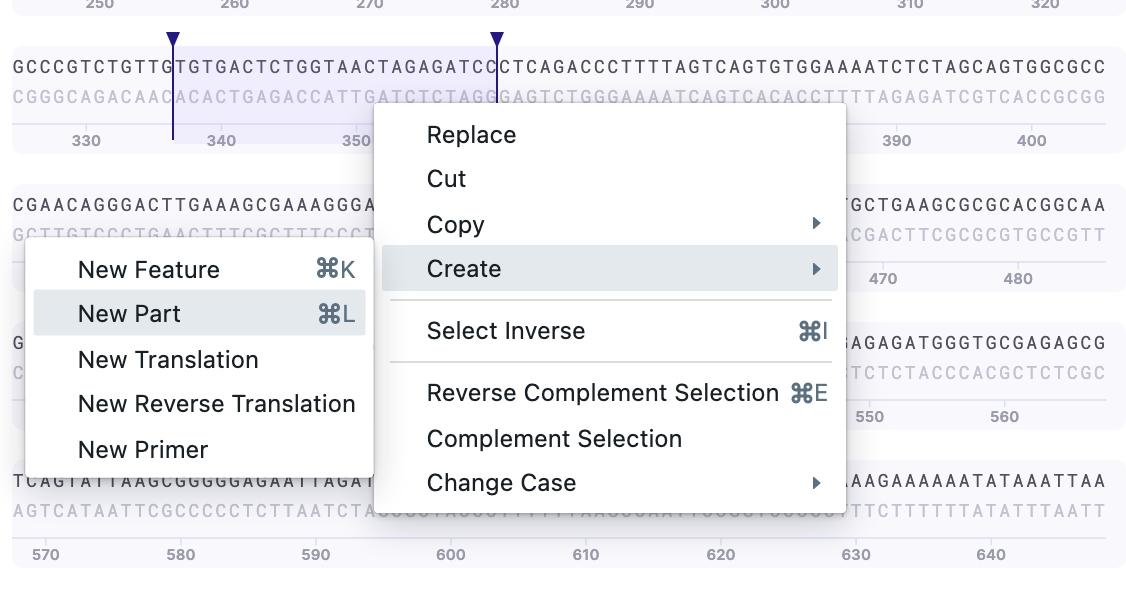
2) Write the name and confirm the strand. You can pick the type of feature as well. You may add several Notes about the selected part of the sequence. Press “Save”. The saved feature will be now displayed.
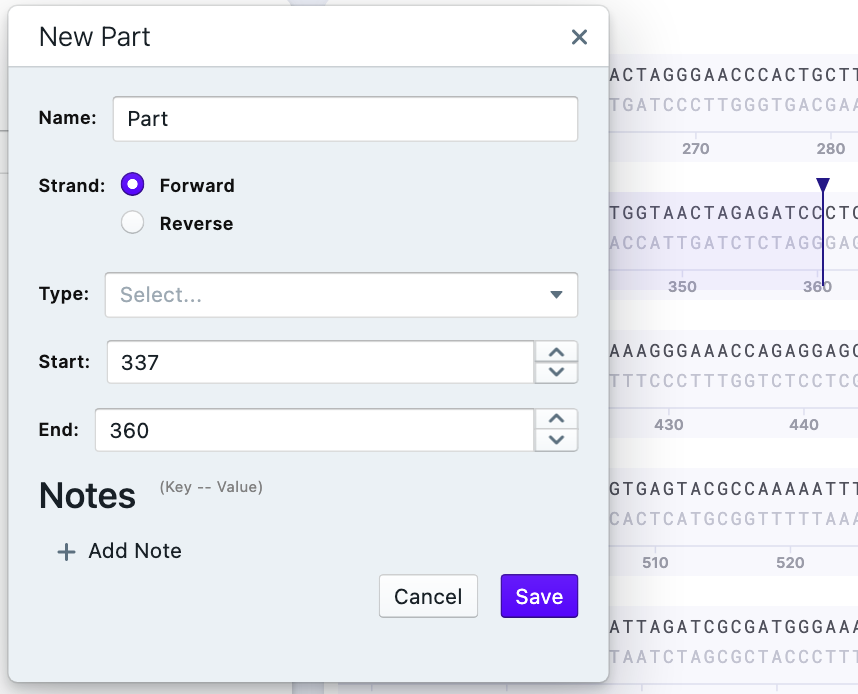

Insert Common Feature
For the convenience of researchers, Mendelgen features a unique option – to Insert Common Feature. We provide a library of the most common sequences that are used to construct vectors. For example, promoters, or antibiotic resistance genes.
1) Move caret to the position where you want your feature to be inserted.
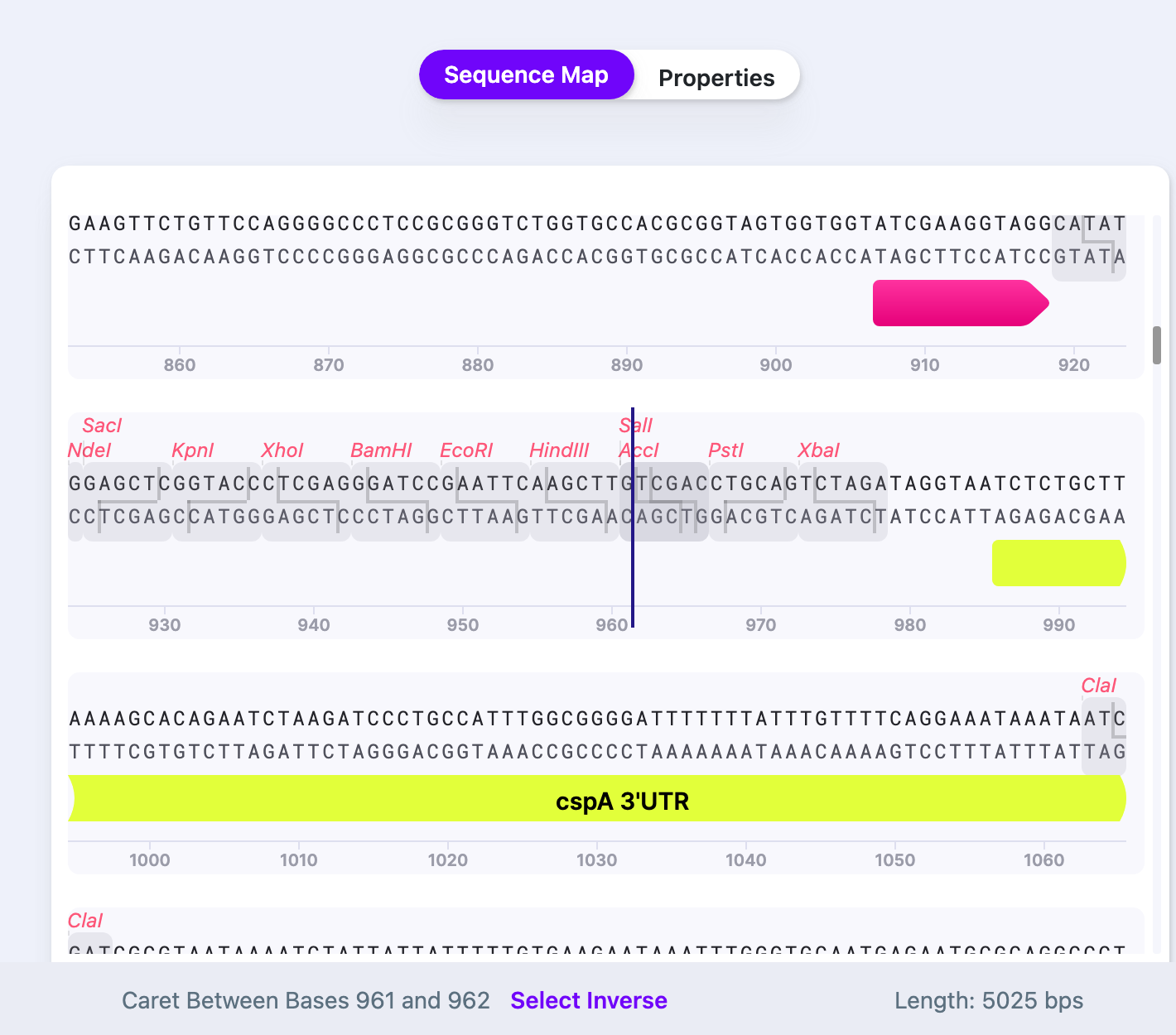
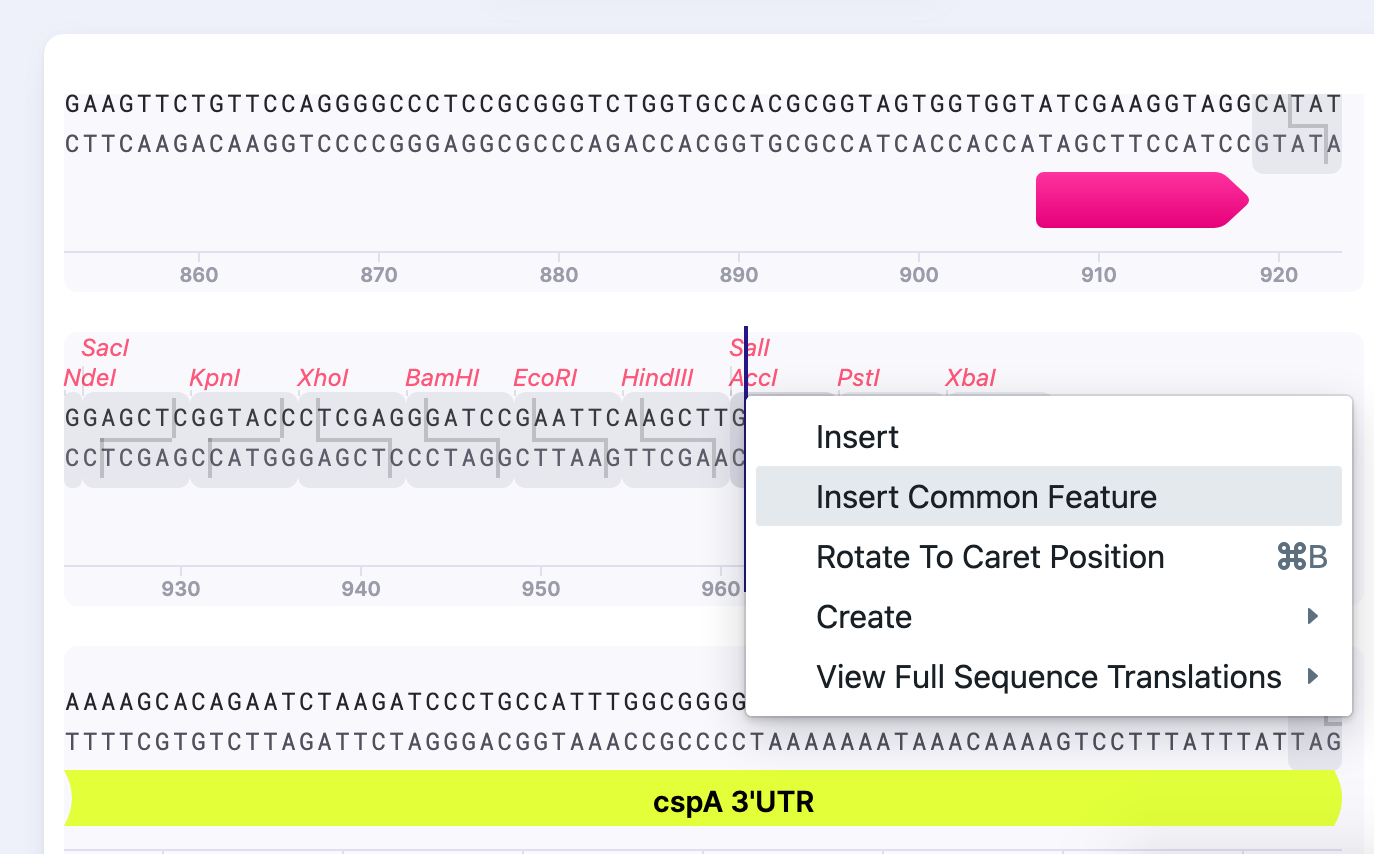
2) Right click and select “Insert Common Feature”.
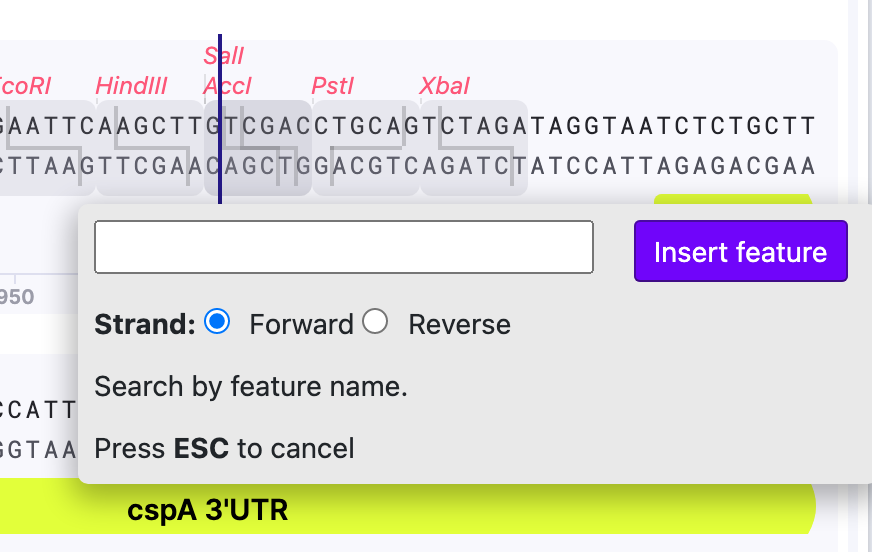
3) In the search window that pops up, you can search feature by name and select the strand.
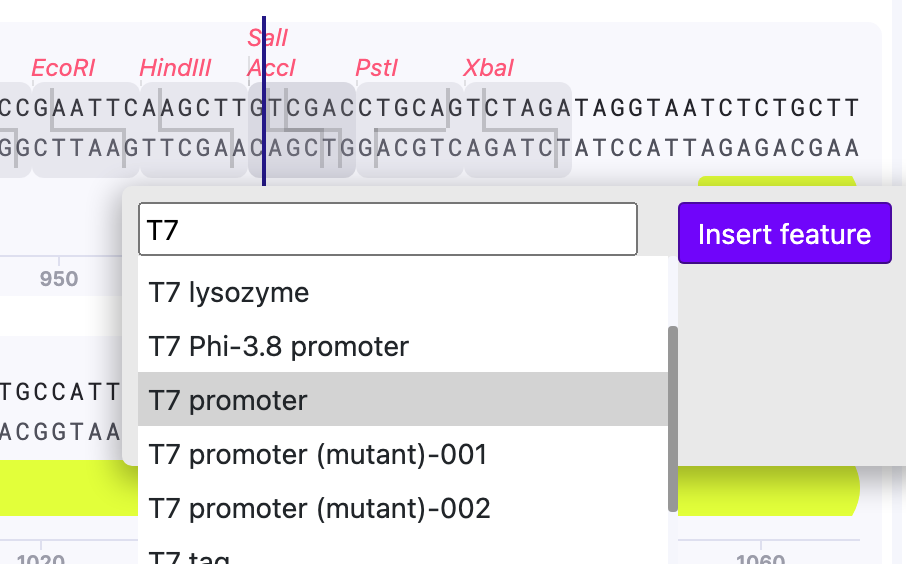
4) After you located the feature, press “Insert Feature”.
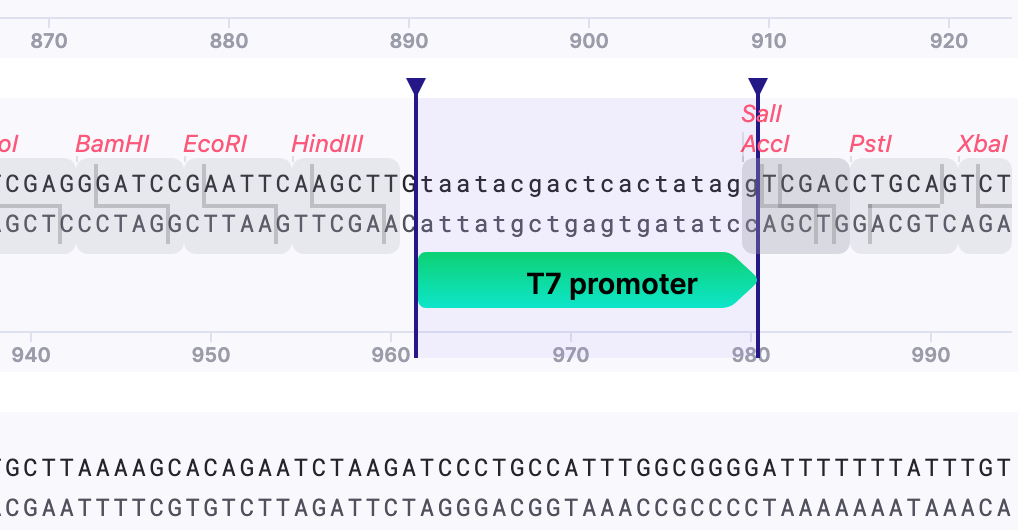
For example, my vector lacks T7 promoter, which is required for cell-free expression. Mendelgen helps to easily enter the sequence.
The sequence is inserted and annotated automatically.
Saving a Sequence
The sequence will be saved in your workspace. To save it on your computer, use the “Export” function.
With hotkey: press ⌘/Ctrl + S or ⌘/Ctrl + Shift + S (Save As).
Manually: press the Save icon on the Control Panel. You have an option to rename your sequence and overwrite the existing one.
Exporting a Sequence
The Export function is available for downloading your sequence as a GenBank file, FASTA file, or JSON file. Press the “Export” button on the control panel and select the file type you would like to download.
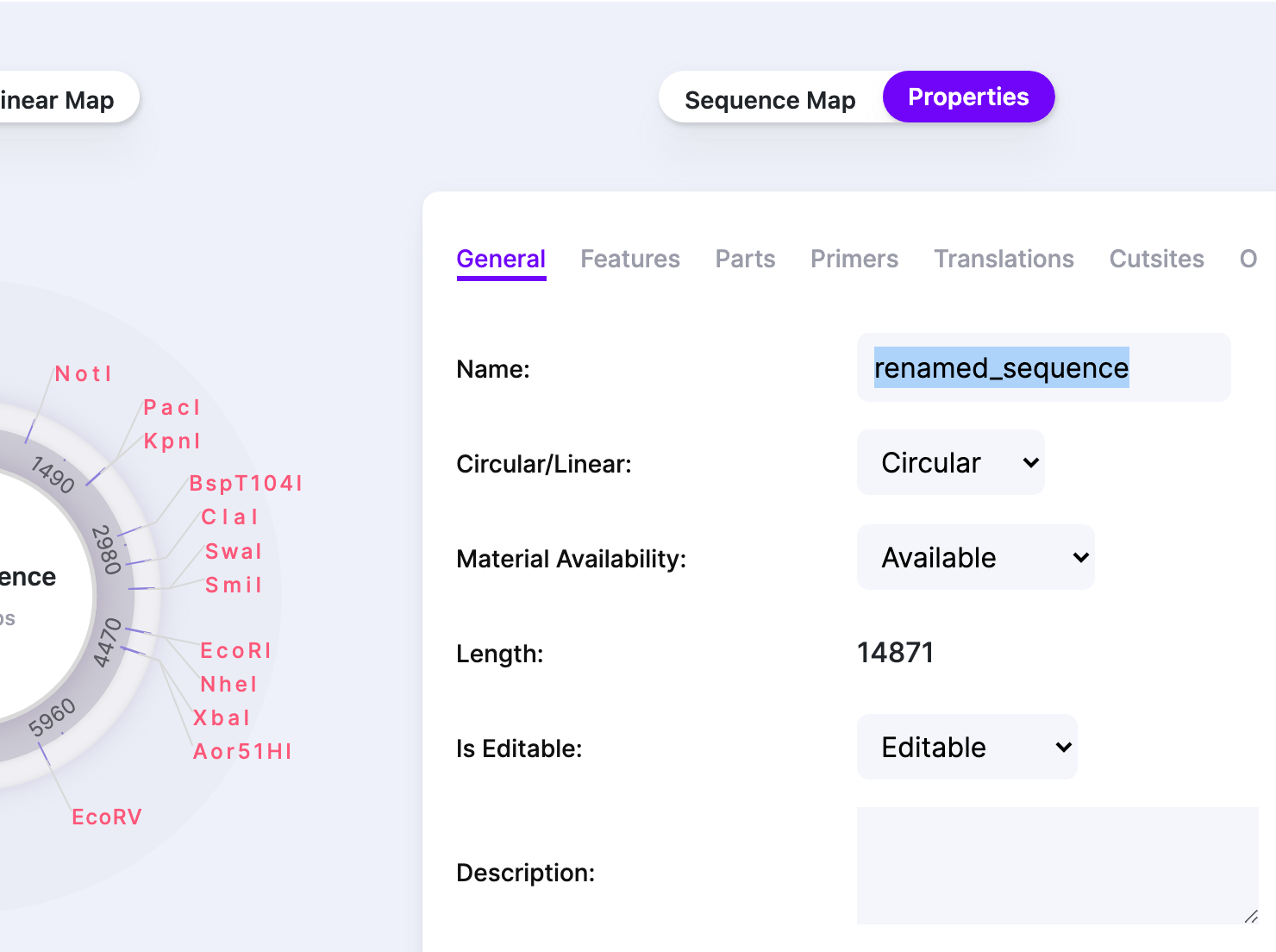
Renaming a Sequence
On the main window, select “Properties” next to “Sequence Map”. In the tab “General”, rename the sequence in the “Name” field.